Dr. Luca Donati
Zuse Insitute Berlin

MATH+ Student Research Group
- Topic Estimating pH-dependent rates by machine learning techniques
- Location Room 046 at Takustr. 9 (FU Berlin) on Friday from 10 to 12
- The seminar will start on 12 May.
-
Molecular Dynamics (MD) simulations are a powerful tool to study the behavior of molecules at the atomic level.
A wide range of computational methods have been developed to analyze MD simulations and to extract meaningful information such as transition rates.
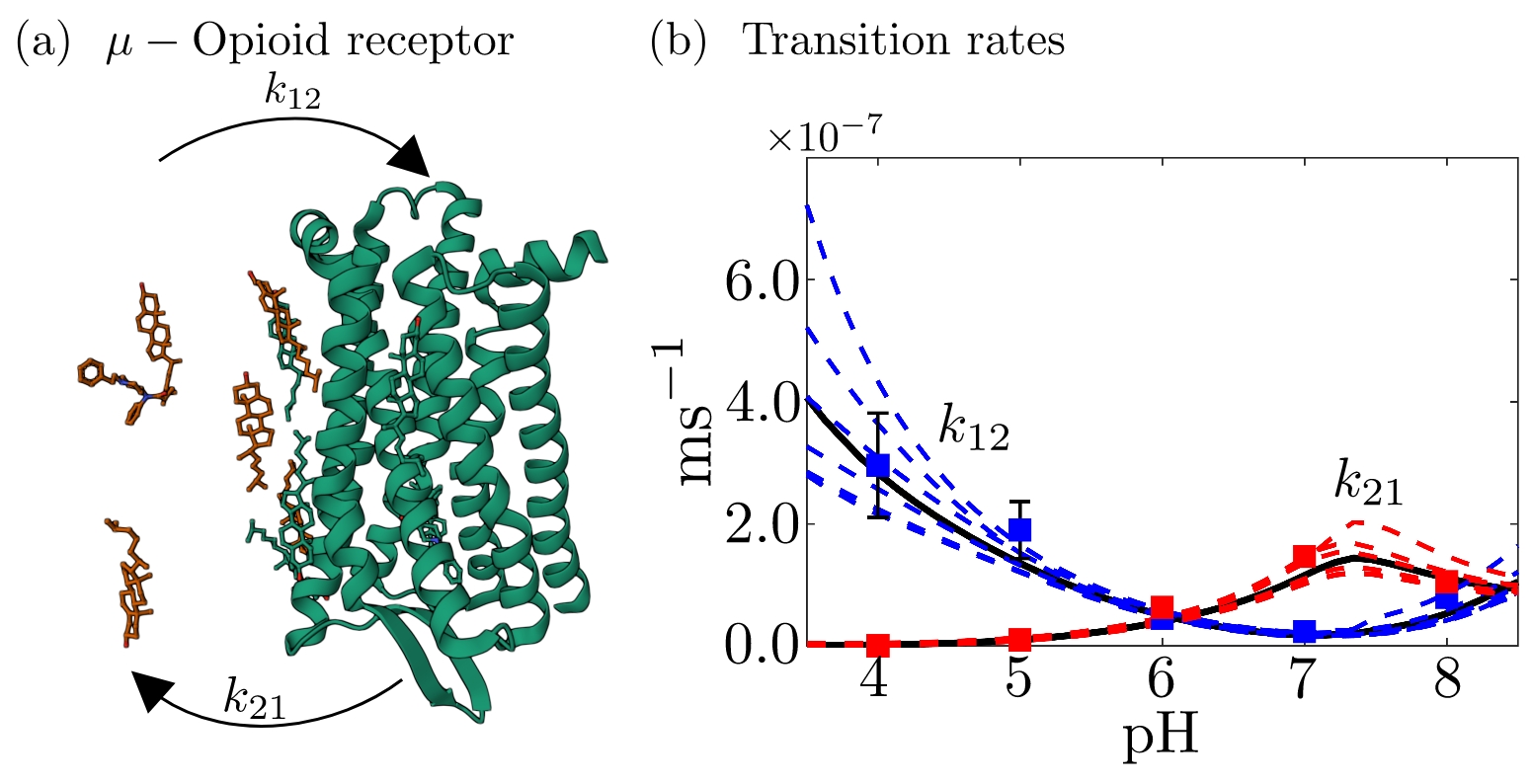
- The purpose of the Student Research Group is to study and extend these methods to determine how transition rates of molecular systems are affected by the pH of the environment [1,2].
- The results of this project will be relevant in studying the μ-opioid receptor system, whose activation and the emergence of adverse effects by opioids depend on the pH of the cell membrane within which it resides [3].
- The project is highly interdisciplinary and students from physics, chemistry, and mathematics are encouraged to participate.
- Please contact me if you are interested in the project.
- References
- L. Donati, and M. Weber. In: J. Chem. Phys. 157.22 (2022), p. 224103, DOI.
- R. J. Rabben, S. Ray, and M. Weber. In: J. Chem. Phys. 153.11 (2020), p. 114109, DOI.
- G. Del Vecchio, et al. In: Sci. Rep. 9 (2019), p. 19344, DOI.
- First lecture
- Introduction
- Simulation of fentanyll and μ-opioid receptor
- Notes on Hamiltonian Dynamics
- Second lecture
- Notes of second lectures
- Introduction to Python
- Jupyter notebook with basic commands (right click, save link as)
- Jupyter notebook with harmonic oscillator 1 (right click, save link as)
- Third lecture
- Notes of third lectures
- Jupyter notebook with harmonic oscillator 2 (right click, save link as)
- Fourth lecture
- Notes about Taylor expansion
- Notes about numerical integrators
- Jupyter notebook with some useful command (right click, save link as)
- Jupyter notebook with Taylor expansion example (right click, save link as)
- Jupyter notebook with complete solution of harmonic oscillator (right click, save link as)
- Fifth lecture
- Notes of fifth lectures
- Jupyter notebook with the solution of the 1D Morse potential (right click, save link as)
- Jupyter notebook with 2D diatomic molecule, complete until force calculation (right click, save link as)
- Sixth lecture
- Jupyter notebook with 2D diatomic molecule, complete (right click, save link as)
- Jupyter notebook with Gaussian distribution 1, complete (right click, save link as)
- Seventh lecture
- Jupyter notebook with Gaussian distribution 2 (right click, save link as)
- Jupyter notebook with Gaussian distribution 3 (right click, save link as)
- Eighth lecture
- Jupyter notebook with Brownian motion (right click, save link as)
- Notes on Brownian motion
- Ninth lecture
- Jupyter notebook with Brownian motion 2 (right click, save link as)
- Jupyter notebook with Langevin dynamics (right click, save link as)
- Tenth lecture
- Jupyter notebook with Python exercises (right click, save link as)
- Jupyter notebook with Langevin dynamics and rate calculation (right click, save link as)

